
The small populations at zero transfer efficiency in D (note the axis scaling and the small amplitudes of this population compared with E) originate from incomplete elimination of molecules with inactive acceptor fluorophores by pulsed interleaved excitation. The vertical lines in C– E indicate the positions of the FRET efficiency peaks of the native population of the respective rhodanese variants. Black lines indicate the DnaK–rhodanese complex population resulting from a fit that takes into account the residual population of refolded and DnaJ-bound rhodanese. ( E) FRET efficiency histograms of DnaK–rhodanese complexes (0.5 µM DnaJ, 10 µM DnaK, and 1 mM ATP DnaK and DnaJ were added simultaneously to rhodanese). ( D) FRET efficiency histograms of DnaJ–rhodanese complexes (0.5 µM DnaJ). ( C) FRET efficiency histograms of native rhodanese (gray) and denatured rhodanese under native conditions transiently populated in the microfluidic mixer (colored, measured 125 ms after dilution of rhodanese into native conditions).
#Chaperone protein code#
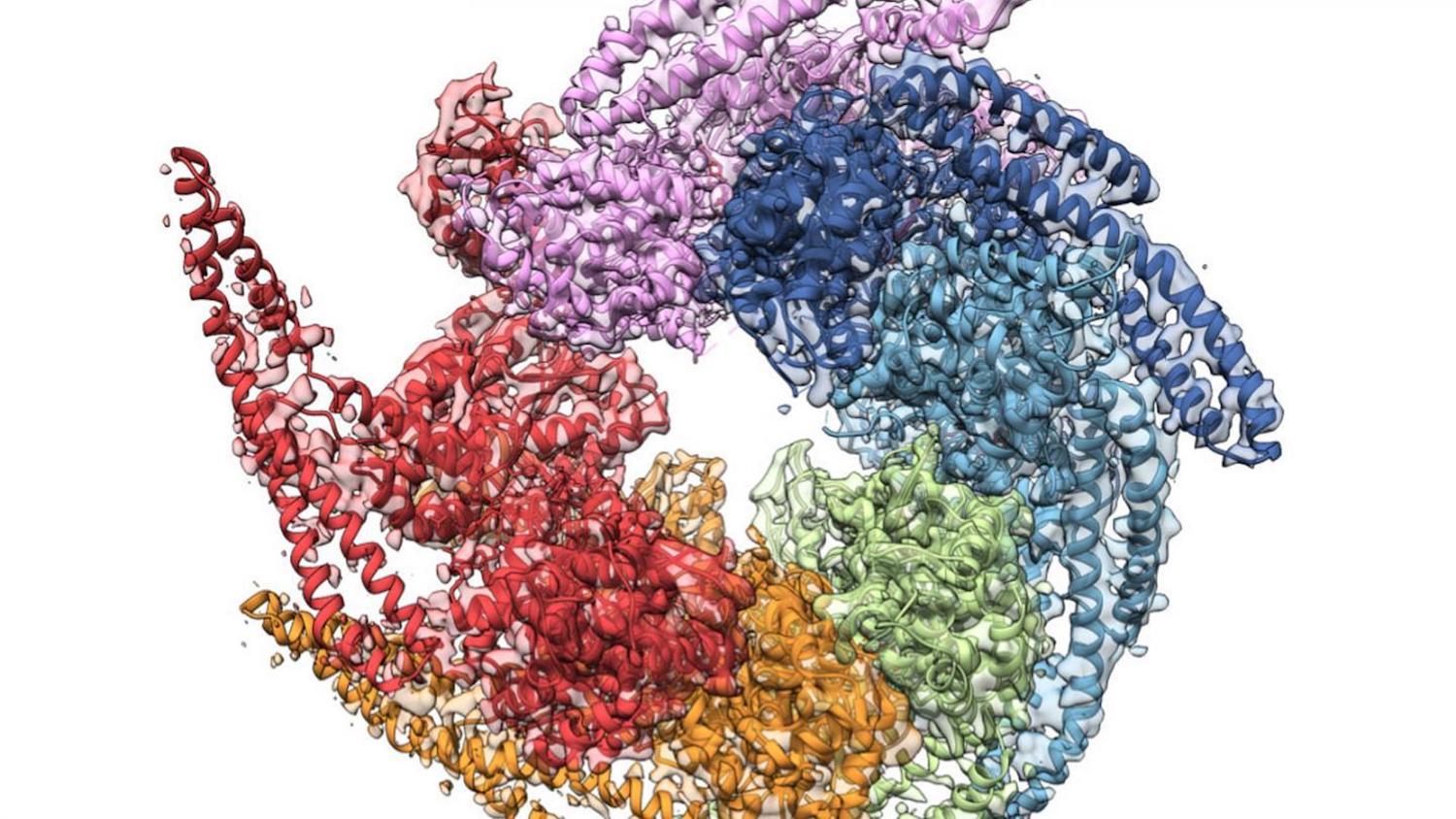
( B) Surface representation of rhodanese (PDB ID code 1RHS) with the subdomains indicated in different gray levels and the label positions of fluorescent dyes for single-molecule FRET measurements shown schematically. ( A) Illustration of the DnaK–ATPase cycle. DnaK expands the denatured substrate protein. To better understand this important link between chaperone action and function, we probed the conformation of a substrate protein along the different stages of the chaperone cycle of DnaK with single-molecule Förster resonance energy transfer (smFRET), correlation spectroscopy, and microfluidic mixing.įig.
#Chaperone protein free#
However, as for other chaperone systems ( 16), surprisingly little is known about how these forces and the resulting constraints of the underlying free energy surfaces affect the conformations of the denatured or misfolded substrate proteins. Since this ATP-driven cycle can even solubilize protein aggregates ( 11, 12), substantial forces must be transduced to the substrate protein ( 13– 15).

Driven by the following GrpE-catalyzed ADP–ATP exchange, the DnaK–substrate complex dissociates ( 10). Substrate and DnaJ synergistically trigger DnaK’s ATPase activity, which leads to locking of the substrate in the DnaK–ADP complex, with DnaK in the closed conformation. Many denatured or misfolded substrate proteins are first captured by DnaJ and subsequently transferred to the DnaK–ATP complex, with DnaK in an open conformation.

The Hsp70 chaperone DnaK from Escherichia coli together with its co-chaperone DnaJ and the nucleotide exchange factor GrpE form an ATP-driven catalytic reaction cycle ( 7) ( Fig. A remarkable example are the heat shock protein (Hsp) 70 chaperones, which are essential in prokaryotes and eukaryotes and are involved in co-translational folding, refolding of misfolded and aggregated proteins, protein translocation, and protein degradation ( 9). For several chaperone systems, these cycles have been investigated in great detail by experiment and simulation ( 6– 8). To assist protein folding, many chaperones proceed through complex conformational cycles in an ATP-dependent manner ( 3– 5). Molecular chaperones have evolved as an essential part of the cellular machinery that facilitates such processes in the complex and crowded environment of a living cell ( 1, 2). Maintaining protein homeostasis in vivo requires a tight regulation of protein folding to prevent misfolding and aggregation. Molecular simulations indicate that hard-core repulsion between the multiple DnaK molecules provides the underlying mechanism for disrupting even strong interactions within the substrate protein and preparing it for processing by downstream chaperone systems. The following ATP-dependent binding of multiple DnaK molecules, however, leads to a surprisingly large expansion of denatured rhodanese. We find that in a first step the ATP-independent binding of DnaJ to denatured rhodanese results in a compact denatured ensemble of the substrate protein. Here we use single-molecule fluorescence spectroscopy to investigate how the DnaJ–DnaK chaperone system alters the conformational distribution of the denatured substrate protein rhodanese. However, understanding the molecular basis of how chaperones prevent such undesirable interactions requires the conformational changes within substrate proteins to be probed during chaperone action. Molecular chaperones are an essential part of the machinery that avoids protein aggregation and misfolding in vivo.


 0 kommentar(er)
0 kommentar(er)
